-Search query
-Search result
Showing all 42 items for (author: mizuno & n)
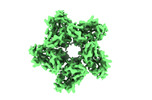
EMDB-35143: 
Cryo-EM structure of the zeaxanthin-bound kin4B8
Method: single particle / : Murakoshi S, Chazan A, Shihoya W, Beja O, Nureki O

PDB-8i2z: 
Cryo-EM structure of the zeaxanthin-bound kin4B8
Method: single particle / : Murakoshi S, Chazan A, Shihoya W, Beja O, Nureki O

EMDB-26794: 
Tomogram of platelet filopodia in presence of SARS-CoV-2 spike protein
Method: electron tomography / : Kuhn CC, Basnet N, Bodakuntla S, Nichols S, Martinez-Sanchez A, Agostini L, Soh YM, Takagi J, Biertumpfel C, Mizuno N

EMDB-26796: 
Tomogram of platelet protrusion with microtubule in presence of SARS-CoV-2 spike protein
Method: electron tomography / : Kuhn CC, Basnet N, Bodakuntla S, Nichols S, Martinez-Sanchez A, Agostini L, Soh YM, Takagi J, Biertumpfel C, Mizuno N

EMDB-26798: 
SARS-CoV-2 spike protein closed conformation
Method: single particle / : Kuhn CC, Basnet N, Bodakuntla S, Alvarez-Brecht P, Nichols S, Martinez-Sanchez A, Agostini L, Soh YM, Takagi J, Biertumpfel C, Mizuno N

EMDB-33293: 
Cryo-EM structure of Cytochrome bo3 from Escherichia coli, apo structure with DMSO
Method: single particle / : Nishida Y, Shigematsu H, Iwamoto T, Takashima S, Shintani Y

EMDB-33294: 
Cryo-EM structure of Cytochrome bo3 from Escherichia coli, the structure complexed with an allosteric inhibitor N4
Method: single particle / : Nishida Y, Shigematsu H, Iwamoto T, Takashima S, Shintani Y

PDB-7xmc: 
Cryo-EM structure of Cytochrome bo3 from Escherichia coli, apo structure with DMSO
Method: single particle / : Nishida Y, Shigematsu H, Iwamoto T, Takashima S, Shintani Y

PDB-7xmd: 
Cryo-EM structure of Cytochrome bo3 from Escherichia coli, the structure complexed with an allosteric inhibitor N4
Method: single particle / : Nishida Y, Shigematsu H, Iwamoto T, Takashima S, Shintani Y

EMDB-25485: 
In situ cryo-electron tomography reveals local cellular machineries for axon branch development (premature branch)
Method: electron tomography / : Nedozralova H, Basnet N, Ibiricu I, Bodakuntla S, Biertumpfel C, Mizuno N

EMDB-25486: 
In situ cryo-electron tomography reveals local cellular machineries for axon branch development (mature branch)
Method: electron tomography / : Nedozralova H, Basnet N, Ibiricu I, Bodakuntla S, Biertumpfel C, Mizuno N

EMDB-12637: 
Cryo-EM structure of human integrin alpha5beta1 in the half-bent conformation
Method: single particle / : Schumacher S, Dedden D, Vazquez Nunez R, Matoba K, Takagi J, Biertumpfel C, Mizuno N

PDB-7nxd: 
Cryo-EM structure of human integrin alpha5beta1 in the half-bent conformation
Method: single particle / : Schumacher S, Dedden D, Vazquez Nunez R, Matoba K, Takagi J, Biertumpfel C, Mizuno N

EMDB-12634: 
Cryo-EM structure of human integrin alpha5beta1 (open form) in complex with fibronectin and TS2/16 Fv-clasp
Method: single particle / : Schumacher S, Dedden D, Vazquez Nunez R, Matoba K, Takagi J, Biertumpfel C, Mizuno N

PDB-7nwl: 
Cryo-EM structure of human integrin alpha5beta1 (open form) in complex with fibronectin and TS2/16 Fv-clasp
Method: single particle / : Schumacher S, Dedden D, Vazquez Nunez R, Matoba K, Takagi J, Biertumpfel C, Mizuno N

EMDB-12160: 
Cilia from MOT7 deletion mutant of Chlamydomonas
Method: subtomogram averaging / : Noga A, Kutomi O, Yamamoto R, Nakagiri Y, Imai H, Obbineni JM, Zimmermann N, Ishikawa T, Wakabayashi K, Kon T, Inaba K

EMDB-12161: 
Chlamydomonas cilia with MOT7-BCCP labeled
Method: subtomogram averaging / : Noga A, Kutomi O, Yamamoto R, Nakagiri Y, Imai H, Obbineni JM, Zimmermann N, Ishikawa T, Wakabayashi K, Kon T, Inaba K

EMDB-12162: 
Microtubule doublet structure from WT Chlamydomonas as a control for MOT7 mutants
Method: subtomogram averaging / : Obbineni JM, Kutomi O, Yamamoto R, Nakagiri Y, Imai H, Noga A, Zimmermann N, Ishikawa T, Wakabayashi K, Kon T, Inaba K

EMDB-10262: 
Yeast 80S ribosome stalled on SDD1 mRNA.
Method: single particle / : Tesina P, Buschauer R, Cheng J, Becker T, Beckmann R

EMDB-10315: 
The cryo-EM structure of SDD1-stalled collided trisome.
Method: single particle / : Tesina P, Buschauer R, Cheng J, Becker T, Beckmann R

PDB-6snt: 
Yeast 80S ribosome stalled on SDD1 mRNA.
Method: single particle / : Tesina P, Buschauer R, Cheng J, Becker T, Beckmann R

PDB-6sv4: 
The cryo-EM structure of SDD1-stalled collided trisome.
Method: single particle / : Tesina P, Buschauer R, Cheng J, Becker T, Beckmann R

EMDB-4772: 
Cryo-EM structure of autoinhibited human talin-1
Method: single particle / : Dedden D, Schumacher S

PDB-6r9t: 
Cryo-EM structure of autoinhibited human talin-1
Method: single particle / : Dedden D, Schumacher S, Zacharias M, Biertumpfel C, Mizuno N

EMDB-4188: 
Cryo-EM structure of microtubule co-polymerized with SSNA1.
Method: helical / : Basnet N, Mizuno N

EMDB-3192: 
Helical reconstruction of amphiphysin N-BAR with a membrane tube radius of 140 Angstrom by cryo-electron microscopy
Method: helical / : Adam J, Basnet N, Mizuno N

EMDB-3193: 
Helical reconstruction of amphiphysin N-BAR with a membrane tube radius of 156 Angstrom by cryo-electron microscopy
Method: helical / : Adam J, Basnet N, Mizuno N

EMDB-3194: 
Helical reconstruction of amphiphysin N-BAR with a membrane tube radius of 131 Angstrom by cryo-electron microscopy
Method: helical / : Adam J, Basnet N, Mizuno N

EMDB-3195: 
Helical reconstruction of amphiphysin N-BAR with a membrane tube radius of 125 Angstrom by cryo-electron microscopy
Method: helical / : Adam J, Basnet N, Mizuno N

EMDB-3196: 
Helical reconstruction of amphiphysin N-BAR with a membrane tube radius of 121 Angstrom by cryo-electron microscopy
Method: helical / : Adam J, Basnet N, Mizuno N

EMDB-2946: 
cryo-EM structure of HET-s prion infectious form
Method: helical / : Mizuno N, Baxa U, Steven AC
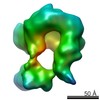
EMDB-2789: 
Cryo-EM map of the Timeless-Tipin-RPA Complex
Method: single particle / : Witosch J, Wolf E, Mizuno N

EMDB-2673: 
Cryo-EM map of CAP-Gly-microtubule complex
Method: helical / : Wang Q, Crevenna AH, Kunze I, Mizuno N

EMDB-2674: 
Cryo-EM map of p150 CAP-Gly-microtubule complex
Method: helical / : Wang Q, Crevenna AH, Kunze I, Mizuno N

EMDB-2675: 
Cryo-EM map of CAP-Gly-microtubule complex
Method: helical / : Wang Q, Crevenna AH, Kunze I, Mizuno N
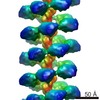
EMDB-2395: 
MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition
Method: helical / : Mizuno N, Dramicanin M, Mizuuchi M, Adam J, Wang Y, Han YW, Yang W, Steven AC, Mizuuchi K, Ramon-Maiques S
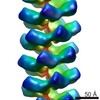
EMDB-2398: 
MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition
Method: helical / : Mizuno N, Dramicanin M, Mizuuchi M, Adam J, Wang Y, Han YW, Yang W, Steven AC, Mizuuchi K, Ramon-Maiques S
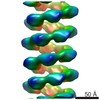
EMDB-2400: 
MuB is an AAA+ ATPase that forms helical filaments to control target selection for DNA transposition
Method: helical / : Mizuno N, Dramicanin M, Mizuuchi M, Adam J, Wang Y, Han YW, Yang W, Steven AC, Mizuuchi K, Ramon-Maiques S

PDB-4bs1: 
MuB is an AAAplus ATPase that forms helical filaments to control target selection for DNA transposition
Method: helical / : Mizuno N, Dramicanin M, Mizuuchi M, Adam J, Wang Y, Han YW, Yang W, Steven AC, Mizuuchi K, Ramon-Maiques S

PDB-4bt0: 
MuB is an AAAplus ATPase that forms helical filaments to control target selection for DNA transposition
Method: single particle / : Mizuno N, Dramicanin M, Mizuuchi M, Adam J, Wang Y, Han YW, Yang W, Steven AC, Mizuuchi K, Ramon-Maiques S

PDB-4bt1: 
MuB is an AAAplus ATPase that forms helical filaments to control target selection for DNA transposition
Method: single particle / : Mizuno N, Dramicanin M, Mizuuchi M, Adam J, Wang Y, Han YW, Yang W, Steven AC, Mizuuchi K, Ramon-Maiques S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
